Our Research Interests
The Godoy lab is interested in investigating the regulation of the bacterial DNA damage responses (DDR) and the mutagenic Y-family DNA polymerases that are part of the DDR. These evolutionarily conserved DNA polymerases are prone to making errors when copying DNA that has been damaged. However, the activity of these DNA polymerases permits cell survival. Our lab uses genetic, molecular, biochemical, and imaging approaches.
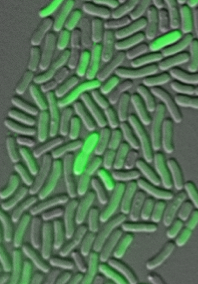
We specifically investigate the regulation of expression of the newly identified DNA damage response in Acinetobacter baumannii, an opportunistic pathogen that is able to quickly gain antibiotic resistances. We have shown that mutagenesis in A. baumannii is dependent on the DDR and thus on the expression of the Y-family DNA polymerases. Our lab has also shown that DDR regulation in A. baumannii is different to that of the Escherichia coli paradigm. Namely, there are two subpopulations formed in response to DNA damage in A. baumannii; one with high expression of the DDR genes and one with low expression (Fig 1). We are currently investigating the molecular mechanisms underlying this novel bimodal regulation of DDR expression.
In addition, we are working to understand the regulation of activity of the Y-family DNA polymerases. We have found that there is a higher order protein complex (Fig 2) composed of one of these Y-family polymerases, DinB, with the equally evolutionarily conserved recombinase RecA and a dimer of the auxiliary protein UmuD. We have evidence suggesting that the protein-protein interactions modulate DinB mutagenesis and access to the replicating fork. We are currently investigating the details of the protein-protein interactions between DinB and RecA.